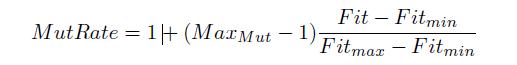| Version 53 (modified by , 14 years ago) (diff) |
|---|
GAIIA
- The team: Mihai Istin - mihai.istin , Andreea Visan - andreea.visan
Abstract
Task scheduling is one of the most important management aspects in a distributed system because this component should achieve two main goals: on the one hand efficient use of available resources and on the other hand high performance in solving tasks. Because of the fact that the scheduling problem is an NP-Complete problem, a near-optimal algorithm is a desired solution. A third goal is the fact that the algorithm should provide the results very fast. This project's aim is to develop a parallel solution for a near-optimal algorithm for dependent task scheduling in distributed systems.
Technologies and Languages
- C/C++
- MPI
- OPENMP
Project Activity (main steps)
- the serial solution - OPENMP tunning - MPI tunning - experimental tests, conclusions
Details
The Problem
The input:
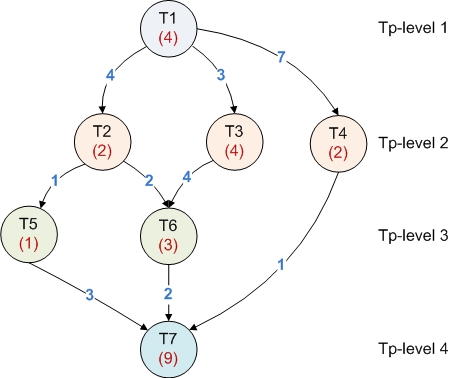
- The graph of tasks that have to be scheduled. Each task is represented as a node while the dependencies between tasks are represented as edges. Each tasks (node) has associated a computational cost and each dependency (edge) has an communication costs.
- The processor topology graph which is also described as a DAG. Each node represent a processor (and the weight represents the speed) while each edge represents the communication cost between processors.
The output:
- The schedule with contains the mapping of tasks on processors and the order of tasks to be executed on the same processor.
The goal:
- load balancing
- minimum makespan
- minimum idle times
The proposed algorithm
Our solution is based on a hybrid evolutionary algorithm with combines the benefits of genetic and immune algorithms algorithms in order to provide efficient solutions. It is known that the convergence time of genetic algorithms is highly influenced by the average fitness of the initial population. So if we could find a way to provide a population with an good average fitness at the initialization phase of the genetic algorithm, the algorithm will converge faster.
Immune algorithms are also bio-inspired algorithms that are based on the human immune system model. They evolve a population of antibodies in order to fit better the antigens that are threatening the immune system.
As a computational model, immune algorithms can be designed to evolve a population of chromosomes representing the antibodies in order to have a good fitness. The implementation of such algorithms is based on the clonal selection principle which model the response of an immune system to an intruder. Basically, the current population of antibodies is evaluated and the best individuals (individuals with the best fitness) are selected for the maturation process. During this process, for each selected individual will be made a number of clones proportionally with its fitness. Then each clone will suffer multiple mutations. The number of mutations will be inverse proportionally with its fitness. Then, the clones are evaluated, and the best are selected in order to survive to the next generation. The antibodies with the lowest fitness will be removed from the current population.
The main advantage of immune algorithms is that they are capable of evolving a population with a good average fitness after only several iteration. On the other hand these algorithms have also a drawback: as a consequence of the fact that the mutation rate varies inverse proportionally with the fitness, the best individuals will be similar. But if we are aware of this drawback we can control the diversity of the evolved population by using only a few generations and by using reinsertion.
The genetic algorithm
1. Chromosome representation
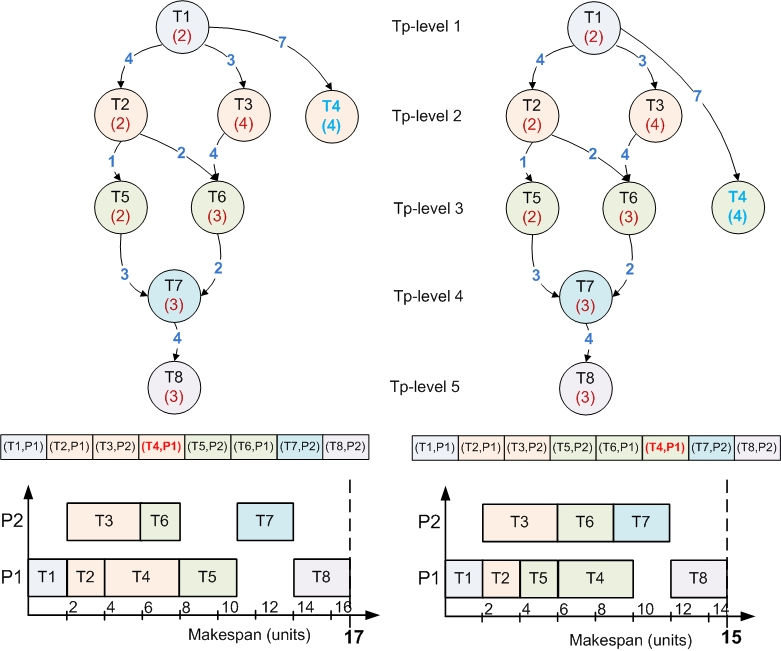
The chromosome representation is based on the topological levels of the nodes. The order of the nodes within a chromosome is a valid topological sort where the first node is an entry node (it has no successors) while the last node is an exit node (it has no successor).
This chromosome encoding assures that none of the genetic operators that will be further described will not violate the dependencies between tasks. So this seems to be a very convenient encoding, but it also has drawbacks which consist in the fact that the explorable solution space will not be entirely covered. This happens because there are some special nodes that can be placed on multiple topological levels without violating the topological order. These nodes are called floating nodes and are characterized as nodes that have a maximum topological level grater than their minimum topological level. The minimum topological level is the level computed during an usual topological sort, while the maximum topological level is the one computed on the transposed graph.
2. Genetic operators
- Single Point Crossover
The crossover operator for a chromosome representation used follows the following steps:
- a cut position is randomly selected ( in the above example the cut point is chosen between genes four and five) and for each chromosome results a head and a tail segment;
- the first offspring is obtained by joining the head segment of the first parent with the combination of the tail segments both parents; more specific, for each gene the task is taken from the first parent and the assigned task is taken from the parent;
- the second offspring is obtain the same way, using the head segment of the first parent and the combination of tail segments of both parents, but this time, the tasks are taken from the second parent while the assigned processors are taken from the first parent
b Simple Gene Mutation
First is selected a chromosome and a randomly selected gene is changed by assigning the task described by the gene to a new processor with the earliest start time.
- Swap Gene Mutation
It is selected a processor that is assigned to run at least two jobs with topological level. Then are randomly selected two tasks with the same topological level assigned to the current processor and the tasks are interchanged if the first task to execute has less children then the second one
- Topological Hyper-Mutation
Whenever a chromosome is affected by this type of mutation, a node from the free node list is randomly selected. Its current topological level is increased and its position in the chromosome is modified in order to reveal the new topological order. Then, all direct predecessors of the selected node are investigated to decide whether they became free floating nodes. If they do, they are inserted in free floating node list corresponding to the current chromosome. If the modified node is no longer a free floating node list (its topological level is equal to the minimum topological level of its successors minus 1), it is removed from the list.
3. Fitness function
The fitness function is an essential element in a GA. It gives an appreciation of the quality of a potential solution according to the problem’s specification. For the scheduling problem, the goal is to obtain task assignments that ensure minimum execution time, maximum processor utilization, a well balanced load across all machines and last but not least to ensure that the precedence of the task is not violated. According to the chromosome encoding and genetic operators presented previously all individuals respect the task DAG, so the focus should be on the other goals of the problem.
For the fitness function we will use the following formula:
4. Selection Method
As a selection method we use a tournament selection algorithms. Basically this algorithm has two steps. First a given number of chromosomes are randomly selected from the population. Next, the selected individual with the best fitness is chosen to reproduce through crossover.
The Immune Algorithms
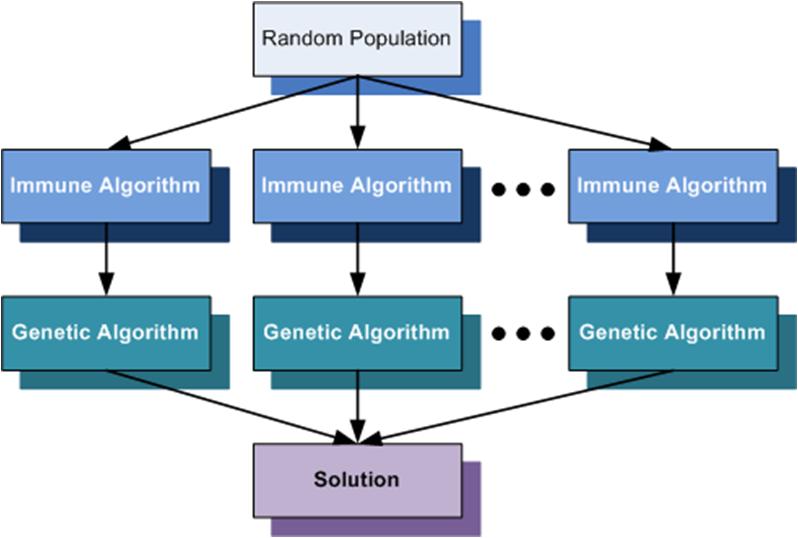
In the following figure is presented the general model for the immune algorithms.
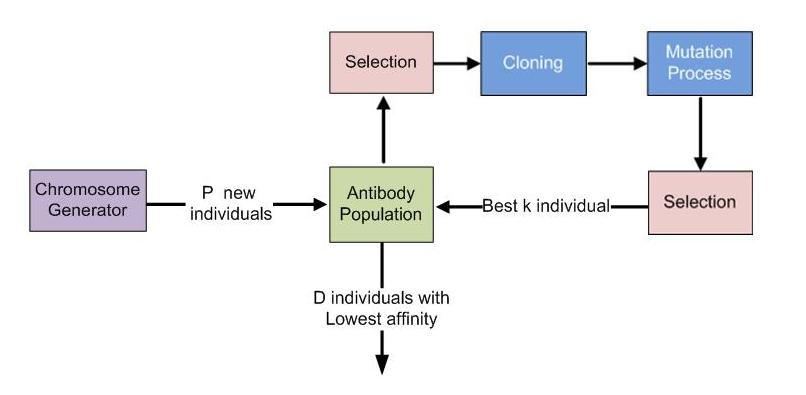
The number of clones is computed using the following formula
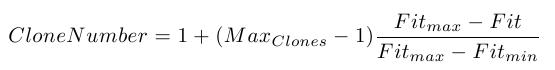
while for the number of mutations we use
The Algorithm Parallelization
1. Serial Solution Analysis
In order to see what are the most computing expensive parts of the designed algorithm we analyzed first the serial version using Sun Studio Performance Analyzer.
In the following figure we can see the time taken by different functions called in the algorithm, sorted in descending order. We highlighted (Red) the functions that consume the most cpu time and that can be implemented using openMP.
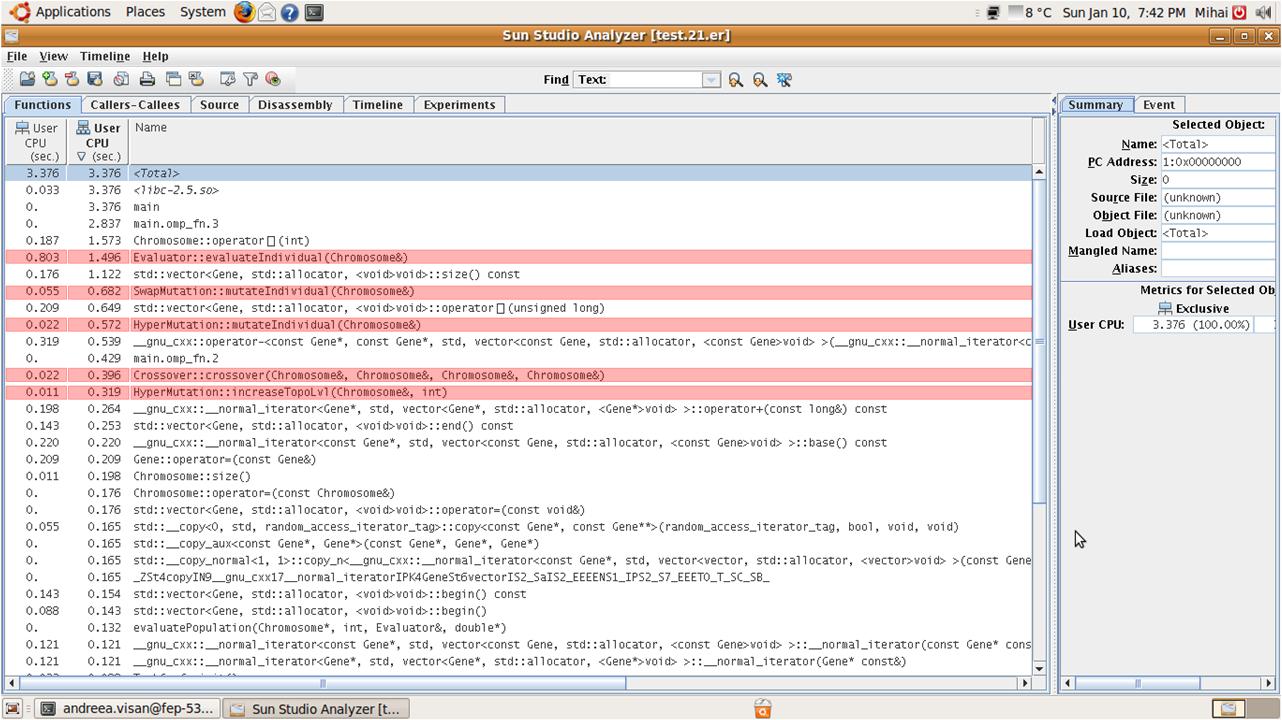
As can be seen in the above figure, most of the functions of the proposed algorithm have significant execution time (10 to 50%) and can be easily parallelized because they are applied independently to each chromosome from the population. In the next paragraphs we will explain what each function does and how it is better to be parallelized.
First we will present the parallel version for the genetic algorithm and then we will present the aspects related to the immune algorithm parallelization.
1.1 The Parallel Genetic Algorithm
The presented analysis reveals that the most time consuming part of the algorithm is the evaluation (or the computation of fitness for each individual). If we analyze the complexity of this phase of the algorithm we will obtain a O(N*M), where N is the number of nodes, M is the maximum number of predecessor that a node can have. More, this function is applied on each of the C chromosomes leading to a total complexity equals to O(C*N*P). If we also consider that this function is applied on each generation, the total complexity is O(G*C*N*M).
Starting from the complexity of the algorithm the easiest and most efficient way to parallelize this function is to apply in on multiple individuals at once. The reason for this is that the evaluation of each individual is independent. This can be done if we use a parallel for with guided scheduling because some individuals survive to the next generation without been affected by mutation and thus there is no need to be evaluated again. After computing the first two terms of the fitness function for each individuals, we also have to compute the last term. This term represents the value of each individual in comparison with the individual with the minimum makespan. For this we have first to compute the minimum makespan. This is also done in parallel. Each thread will compute the minimum makespan from its corresponding chunk (from a static scheduling) and will place the result in a shared memory. Next, the master will compute the overall minimum am will place it in the shared memory. Then, each thread will finally compute the fitness for each individual from its chunk.
For the crossover is used a static scheduling because each two individuals will recombine in order to produce offspring. On the other hand for the mutation operator we will also use a guided scheduling because for the genetic algorithm an individual will suffer a mutation only with a certain probability, so some individuals will not be affected. For the immune algorithm, the proper scheduling strategy is static because each individual will be affected by mutation.
Another interesting approach is used for the selection algorithm. Here the parallelization is not strait forward. The population of chromosomes will be divided in a number of chunks equal to the number of threads. At each step (of the total TOUR_SIZE steps), each thread will randomly choose an unselected individual from a single chunk. The chunks assignment to threads is done in a round robin manner. After each thread has completed the tour, the individuals with the best fitness is selected to survive in the next generation and the process is started over.
1.2 The Parallel Immune Algorithm
First of all we have to mention that for the evaluation of the chromosome we use the same approach presented for the genetic algorithm.
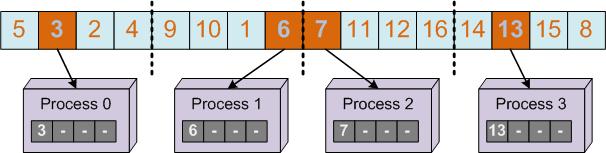
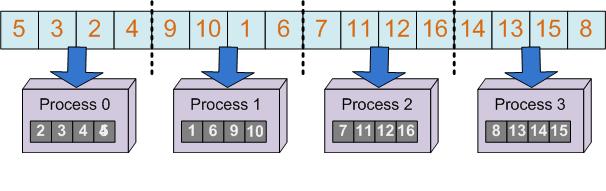
The selection method for the immune algorithm is a simple selection of the best individuals and it is based on choosing the first individuals after they have been sorted. So we will need a parallel sort algorithm. The chosen algorithm is a hybrid algorithm between Quick Sort and Merge Sort. First the population is divided into a number of chunks equal to the number of threads and each thread will sort its chunk using the Quick Sort algorithm (first figure). In the second phase each two distinct chunks will be merged until we will obtain a sorted vector (second figure).
In order to produce the population of clones we have first to determine the number of clones corresponding to each individual. Then we have to distribute the cloning process in order to assure load balancing among threads. For this we use a parallel algorithm to compute the partial sums. The same approach will also be used for the mutation process.
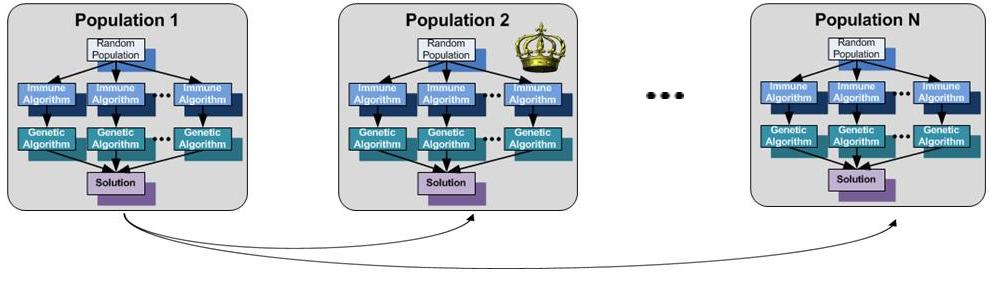
1.3 The Parallel Execution Model
- The Distributed Execution Model
Compiling, Running, Profiling
- the OpenMP version
g++ -g -fopenmp -o scheduler main.cpp ...
qsub -q ibm-quad.q -pe openmpi 1 -v OMP_NUM_THREADS=4 script.h
Experimental results
In the following figures, are represented the results obtained using Sun Studio Performance Analyzer. The input tasks graph contains 500 nodes. The experimental tests were made only for the OpenMP version of the project for different numbers of threads.
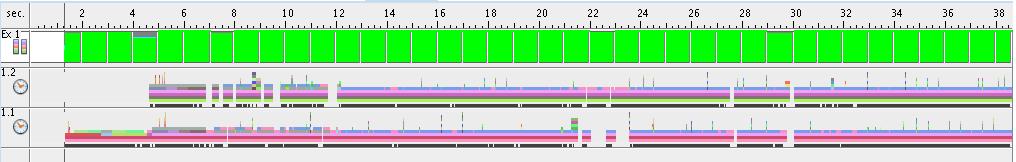
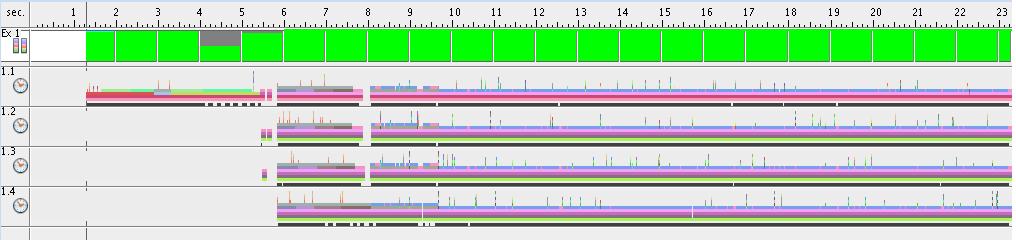
- Results for 1 thread
- Results for 2 threads
- Results for 4 threads
- Results for 8 threads
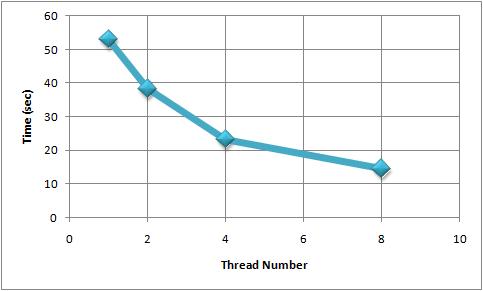
- Evolution - Summary
The next figure summarizes the execution times obtained in the situation of 1, 2, 4 and 8 threads for the experimental tests presented above. As we can observe, the execution time is decreasing with the increasing of the threads number.
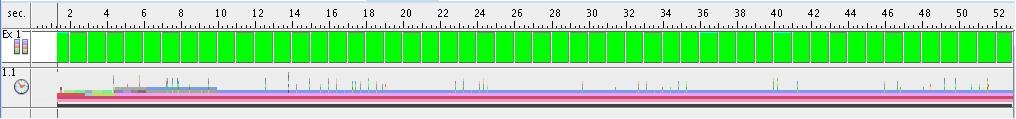
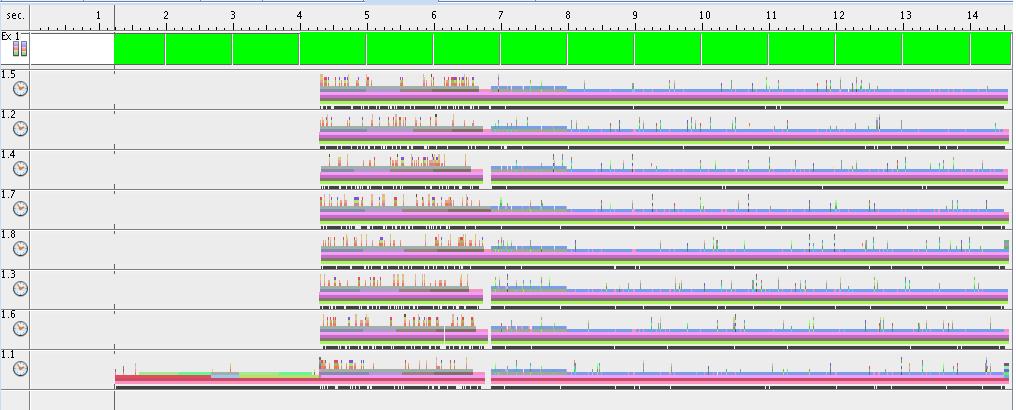
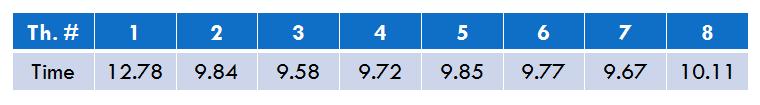
- Execution time for each thread (8 threads situation)
The next figure presents the execution time of each thread in the situation of 8 threads-execution. Thread #1 has a longer execution time because of the fact that it has to do data initialization. All the other threads have a balanced execution time.
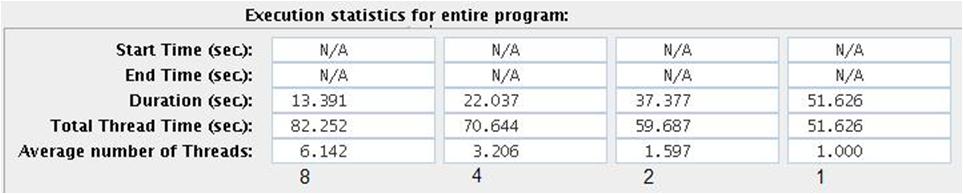
- Execution statistics for the entire program for 1,2,4 and 8 threads
Attachments (23)
-
chromosome.jpg (18.2 KB) - added by 14 years ago.
The chromosome's representation
-
DAG.jpg (52.1 KB) - added by 14 years ago.
The DAG of tasks
-
floating_nodes.jpg (170.8 KB) - added by 14 years ago.
Example of floating nodes
- 1.jpg (29.2 KB) - added by 14 years ago.
-
2.jpg (37.1 KB) - added by 14 years ago.
Execution - 2 threads
-
4.jpg (50.1 KB) - added by 14 years ago.
Execution - 4 threads
-
8.jpg (76.4 KB) - added by 14 years ago.
Execution - 8 threads
-
overview.jpg (124.1 KB) - added by 14 years ago.
Sun Studio Performance Analyzer - function overview
-
ia.jpg (22.6 KB) - added by 14 years ago.
The Immune Algorithm
-
execp.jpg (37.0 KB) - added by 14 years ago.
(Parallel) Execution model
-
execd.jpg (41.3 KB) - added by 14 years ago.
(Distributed) Execution model
- grafic2.jpg (170.1 KB) - added by 14 years ago.
-
grafic1.jpg (16.6 KB) - added by 14 years ago.
Evolution (thread)
- runtime.jpg (21.1 KB) - added by 14 years ago.
- tab.jpg (13.1 KB) - added by 14 years ago.
- tab2.jpg (29.3 KB) - added by 14 years ago.
- form1.jpg (5.8 KB) - added by 14 years ago.
- form2.jpg (6.4 KB) - added by 14 years ago.
- form3.jpg (9.6 KB) - added by 14 years ago.
- tour.jpg (19.7 KB) - added by 14 years ago.
- sort1.jpg (21.4 KB) - added by 14 years ago.
- sort2.jpg (16.6 KB) - added by 14 years ago.
- multi.png (25.4 KB) - added by 14 years ago.
Download all attachments as: .zip
![(please configure the [header_logo] section in trac.ini)](/trac/PP2009/chrome/site/your_project_logo.png)